-Search query
-Search result
Showing 1 - 50 of 61 items for (author: qi & jx)
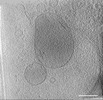
EMDB-39012: 
Representative tomogram of primary glioblastoma stem cell with circular inter-mitochondrial junctions.
Method: electron tomography / : Wang R, Lei H, Wang HX, Qi L, Liu YE, Liu YH, Shi YF, Chen JX, Shen QT
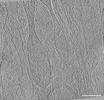

EMDB-39015: 
Representative tomogram of microglia cell with nanotunnel-like structures resembling mitochondrial fission.
Method: electron tomography / : Wang R, Lei H, Wang HX, Qi L, Liu YE, Liu YH, Shi YF, Chen JX, Shen QT
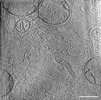
EMDB-39019: 
Representative tomogram of glioblastoma cell with nanotunnel-like structure and inter-mitochondrial junction.
Method: electron tomography / : Wang R, Lei H, Wang HX, Qi L, Liu YE, Liu YH, Shi YF, Chen JX, Shen QT

EMDB-39021: 
Representative tomogram of normal human astrocyte with nanotunnel-like structure which is an extension of the mitochondrial outer membrane.
Method: electron tomography / : Wang R, Lei H, Wang HX, Qi L, Liu YE, Liu YH, Shi YF, Chen JX, Shen QT
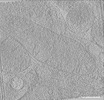

EMDB-39023: 
Representative tomogram of primary glioblastoma differentiated cell with parallel inter-mitochondrial junction.
Method: electron tomography / : Wang R, Lei H, Wang HX, Qi L, Liu YE, Liu YH, Shi YF, Chen JX, Shen QT
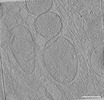

EMDB-39024: 
Representative tomogram of primary glioblastoma stem cell with clustered mitochondria bearing various long-short axis ratios.
Method: electron tomography / : Wang R, Lei H, Wang HX, Qi L, Liu YE, Liu YH, Shi YF, Chen JX, Shen QT

EMDB-36057: 
Cryo-EM map of DAM (a newly designed immunogen for monkeypox virus) in complex with half of the antigen-binding fragment (Fab) of A27D7 and the single-chain fragment variable (scFv) of 7D11
Method: single particle / : Wang H, Yin P, Zheng TT, Han P, Qi JX

EMDB-36056: 
Cryo-EM map of half of DAM (the A35 part) in complex with the antigen-binding fragment (Fab) of A27D7
Method: single particle / : Wang H, Yin P, Zheng TT, Qin LJ, Han P, Qi JX

EMDB-34727: 
Cryo-EM structure of SARS-CoV-2 Omicron BA.1 spike protein in complex with white-tailed deer ACE2
Method: single particle / : Han P, Meng YM, Qi JX

EMDB-34728: 
SARS-CoV-2 Omicron BA.1 spike protein receptor-binding domain in complex with white-tailed deer ACE2
Method: single particle / : Han P, Meng YM, Qi JX

EMDB-34729: 
Cryo-EM structure of SARS-CoV-2 prototype spike protein in complex with white-tailed deer ACE2
Method: single particle / : Han P, Meng YM, Qi JX

EMDB-34730: 
Cryo-EM structure of SARS-CoV-2 prototype spike protein receptor-binding domain in complex with white-tailed deer ACE2
Method: single particle / : Han P, Meng YM, Qi JX

EMDB-35426: 
Cryo-EM structure of SARS-CoV-2 Omicron BA.4/5 spike protein in complex with white-tailed deer ACE2
Method: single particle / : Han P, Meng YM, Qi JX

EMDB-35427: 
Cryo-EM structure of SARS-CoV-2 Omicron BA.4/5 spike protein receptor-binding domain in complex with white-tailed deer ACE2
Method: single particle / : Han P, Meng YM, Qi JX
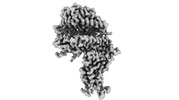
EMDB-33841: 
Cryo-EM structure of SARS-CoV-2 Omicron BA.2.12.1 RBD in complex with human ACE2 (local refinement)
Method: single particle / : Zhao ZN, Xie YF, Qi JX, Gao GF
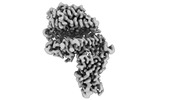
EMDB-33870: 
Cryo-EM structure of SARS-CoV-2 Omicron BA.2 RBD in complex with human ACE2 (local refinement)
Method: single particle / : Zhao ZN, Xie YF, Qi JX, Gao GF
EMDB-34120: 
Cryo-EM structure of SARS-CoV-2 Omicron BA.2 RBD in complex with golden hamster ACE2 (local refinement)
Method: single particle / : Zhao ZN, Xie YF, Chai Y, Qi JX, Gao GF

EMDB-34138: 
Cryo-EM structure of SARS-CoV-2 Omicron BA.2 RBD in complex with mouse ACE2 (local refinement)
Method: single particle / : Zhao ZN, Xie YF, Chai Y, Qi JX, Gao GF

EMDB-34217: 
Cryo-EM structure of SARS-CoV-2 Omicron BA.2 RBD in complex with rat ACE2 (local refinement)
Method: single particle / : Zhao ZN, Xie YF, Chai Y, Qi JX, Gao GF

EMDB-34409: 
Cryo-EM structure of SARS-CoV-2 Omicron BA.4/5 RBD in complex with human ACE2 (local refinement)
Method: single particle / : Zhao ZN, Xie YF, Qi JX, Gao GF

EMDB-34494: 
Cryo-EM map of SARS-CoV-2 Omicron BA.2 spike trimer in complex with human ACE2 (three-RBD-up conformation)
Method: single particle / : Zhao ZN, Xie YF, Qi JX, Gao GF

EMDB-34498: 
Cryo-EM map of SARS-CoV-2 Omicron BA.2.12.1 spike trimer in complex with human ACE2 (three-RBD-up conformation)
Method: single particle / : Zhao ZN, Xie YF, Qi JX, Gao GF

EMDB-34499: 
Cryo-EM map of SARS-CoV-2 Omicron BA.2 spike protein in complex with mouse ACE2
Method: single particle / : Zhao ZN, Xie YF, Chai Y, Qi JX, Gao GF

EMDB-34506: 
Cryo-EM map of SARS-CoV-2 Omicron BA.2 spike trimer in complex with rat ACE2
Method: single particle / : Zhao ZN, Xie YF, Chai Y, Qi JX, Gao GF

EMDB-34509: 
Cryo-EM map of SARS-CoV-2 Omicron BA.4/5 (N658S) spike trimer in complex with human ACE2 (three-RBD-up conformation)
Method: single particle / : Zhao ZN, Xie YF, Qi JX, Gao GF
EMDB-34510: 
Cryo-EM map of SARS-CoV-2 Omicron BA.2 spike trimer in complex with golden hamster ACE2
Method: single particle / : Zhao ZN, Xie YF, Chai Y, Qi JX, Gao GF

EMDB-34227: 
Cryo-EM structure of Singapore Grouper Iridovirus capsid block 1
Method: single particle / : Zhao ZN, Liu CC, Zhu DJ, Qi JX, Zhang XZ, Gao GF

EMDB-34229: 
Cryo-EM structure of Singapore Grouper Iridovirus capsid block 4
Method: single particle / : Zhao ZN, Liu CC, Zhu DJ, Qi JX, Zhang XZ, Gao GF

EMDB-34230: 
Cryo-EM structure of Singapore Grouper Iridovirus capsid block 2
Method: single particle / : Zhao ZN, Liu CC, Zhu DJ, Qi JX, Zhang XZ, Gao GF

EMDB-34235: 
Cryo-EM structure of Singapore Grouper Iridovirus capsid block 3
Method: single particle / : Zhao ZN, Liu CC, Zhu DJ, Qi JX, Zhang XZ, Gao GF

EMDB-34236: 
Cryo-EM structure of Singapore Grouper Iridovirus capsid block 5
Method: single particle / : Zhao ZN, Liu CC, Zhu DJ, Qi JX, Zhang XZ, Gao GF
EMDB-34251: 
Singapore Grouper Iridovirus
Method: single particle / : Zhao ZN, Liu CC, Zhu DJ, Qi JX, Zhang XZ, Gao GF

EMDB-34815: 
One asymmetric unit of Singapore grouper iridovirus capsid
Method: single particle / : Zhao ZN, Liu CC, Zhu DJ, Qi JX, Zhang XZ, Gao GF

EMDB-34731: 
The structure of MPXV polymerase holoenzyme in replicating state
Method: single particle / : Peng Q, Xie YF, Kuai L, Wang H, Qi JX, Gao F, Shi Y

EMDB-33697: 
Cryo-EM structure of SARS-CoV-2 Omicron spike protein (S-6P-RRAR) in complex with human ACE2 ectodomain (one-RBD-up state)
Method: single particle / : Gao GF, Qi JX, Liu S, Zhao ZN

EMDB-33698: 
Cryo-EM structure of hACE2-bound SARS-CoV-2 Omicron spike protein with L371S, P373S and F375S mutations (S-6P-RRAR)
Method: single particle / : Zhao ZN, Xie YF, Qi JX, Gao GF

EMDB-33124: 
Cryo-EM structure of SARS-CoV-2 Omicron spike protein (S-6P-RRAR) in complex with S309 fab
Method: single particle / : Gao GF, Qi JX, Zhao ZN, Liu S, Xie YF

EMDB-33690: 
Cryo-EM structure of apo SARS-CoV-2 Omicron spike protein (S-2P-GSAS)
Method: single particle / : Zhao ZN, Xie YF, Qi JX, Gao GF

EMDB-33699: 
Cryo-EM structure of hACE2-bound SARS-CoV-2 Omicron spike protein with L371S, P373S and F375S mutations (local refinement)
Method: single particle / : Zhao ZN, Xie YF, Qi JX, Gao GF

EMDB-33709: 
Cryo-EM structure of S309-RBD-RBD-S309 in the S309-bound Omicron spike protein (local refinement)
Method: single particle / : Zhao ZN, Xie YF, Qi JX, Gao F

EMDB-33120: 
Cryo-EM structure of SARS-CoV-2 Omicron spike protein (S-6P-RRAR) in complex with human ACE2 ectodomain (two-RBD-up state)
Method: single particle / : Gao GF, Qi JX, Zhao ZN, Liu S, Xie YF

EMDB-33121: 
Cryo-EM structure of SARS-CoV-2 Omicron RBD in complex with human ACE2 ectodomain (local refinement)
Method: single particle / : Gao GF, Qi JX, Zhao ZN, Liu S, Xie YF

EMDB-33123: 
Cryo-EM structure of SARS-CoV-2 Omicron RBD in complex with S309 fab (local refinement)
Method: single particle / : Gao GF, Qi JX, Zhao ZN, Xie YF, Liu S

EMDB-32358: 
Structure of SARS-CoV-2 spike receptor-binding domain Y453F mutation complexed with American mink ACE2
Method: single particle / : Su C, Qi JX, Gao GF

EMDB-32379: 
Structure of SARS-CoV-2 spike receptor-binding domain F486L mutation complexed with American mink ACE2
Method: single particle / : Su C, Qi JX, Gao GF

EMDB-30903: 
Structure of TRPC3 at 2.7 angstrom in high calcium state
Method: single particle / : Chen L, Guo W

EMDB-30904: 
Structure of TRPC3 at 3.06 angstrom in low calcium state
Method: single particle / : Chen L, Guo W

EMDB-30906: 
Structure of TRPC3 gain of function mutation R803C at 3.2 angstrom in 1340nM free calcium state
Method: single particle / : Chen L, Guo W

EMDB-30907: 
Structure of BTDM-bound human TRPC6 nanodisc at 2.9 angstrom in high calcium state
Method: single particle / : Chen L, Guo W

EMDB-30908: 
Structure of SAR7334-bound TRPC6 at 2.9 angstrom
Method: single particle / : Chen L, Guo W
Pages:
 Movie
Movie Controller
Controller Structure viewers
Structure viewers About EMN search
About EMN search



 wwPDB to switch to version 3 of the EMDB data model
wwPDB to switch to version 3 of the EMDB data model
